Functional Genomics and Network Biology
Characterizing the ell-type-specific molecular perturbations potentially driving dysregulation of biological processes in Alzheimer's Disease.
1. Unravelling cell-type-specific molecular network perturbations driving dysregulation of biological processes in AD. (Adeoye et al., 2024).
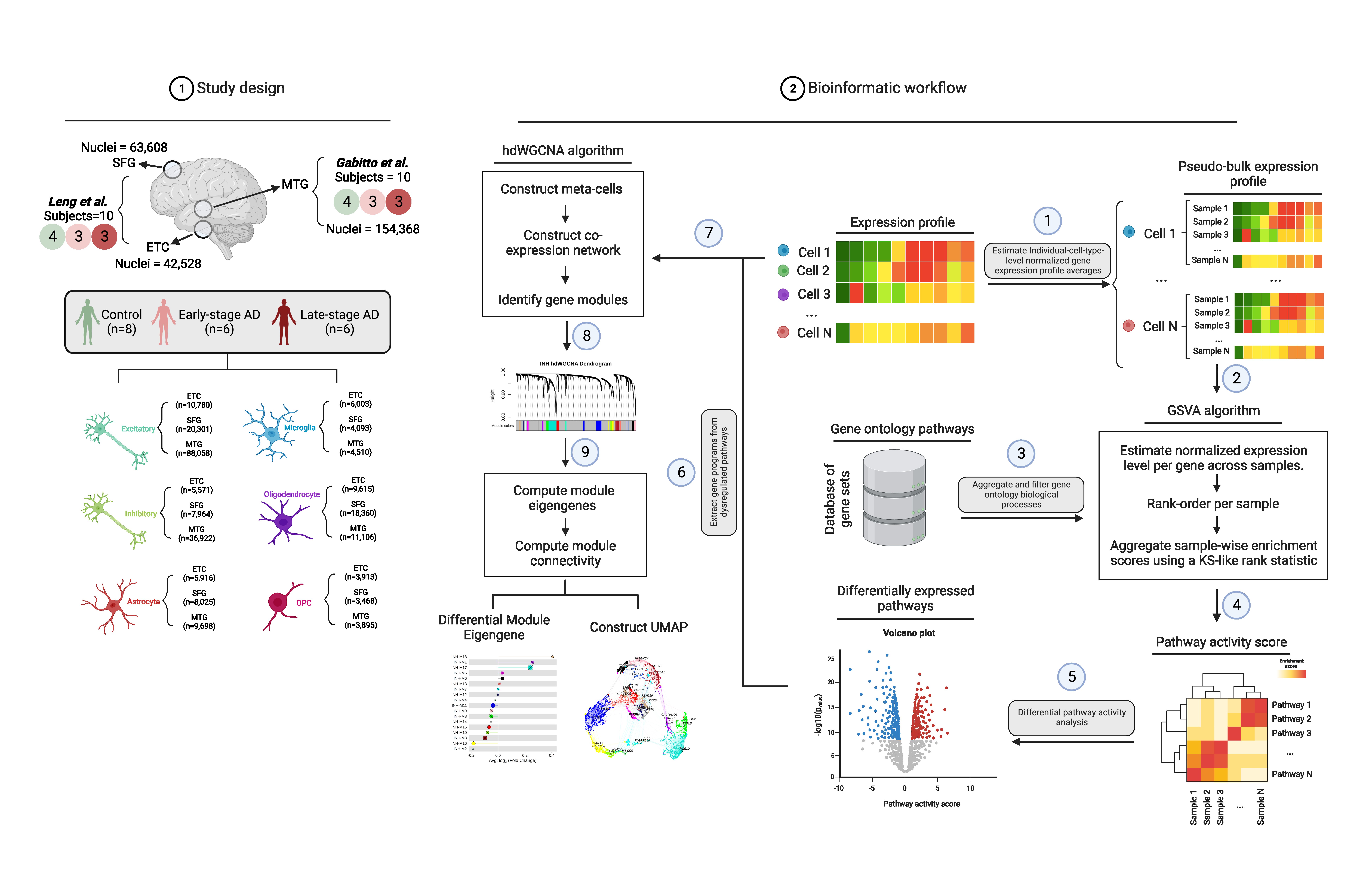
The motivation for this work stems from the recognition of AD as a complex systems disease, where pathology extends beyond isolated molecular changes to encompass intricate interactions within gene networks. While transcriptomic profiling has enabled investigation of cell-type-specific mechanisms in AD, traditional differential gene expression (DEG) analysis fails to capture the full spectrum of disrupted processes underlying the disease. Our work addresses this gap by developing a systems biology approach to uncover the cell-type-specific drivers of biological process perturbations in AD.
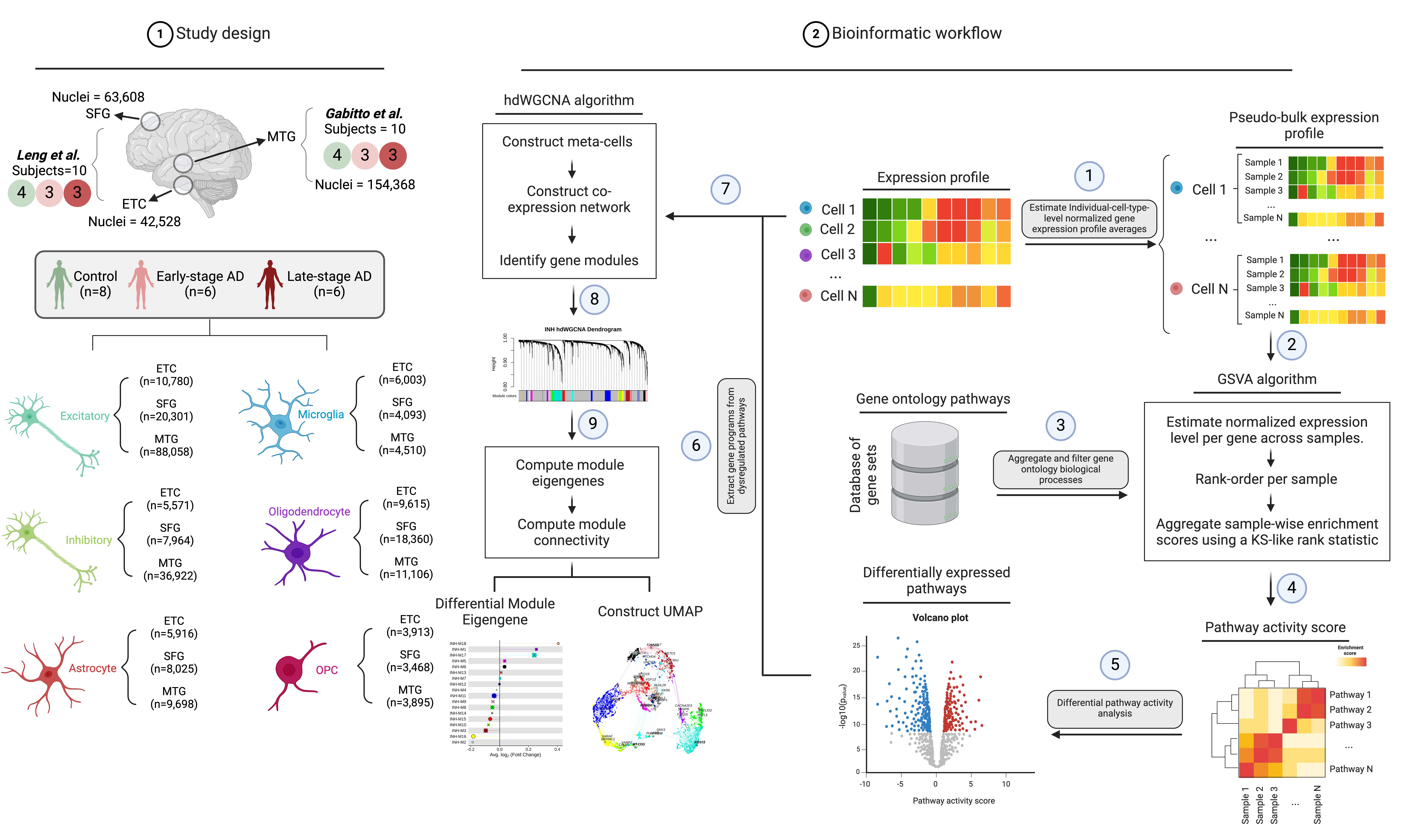
We utilized snRNA-seq data from key AD brain regions and combined differential activity analysis of Gene Ontology biological processes with high-dimensional weighted gene co-expression analysis (hdWGCNA) to provide insights into the functional coherence among genes in perturbed processes. We aggregated gene expression values within each GO process into pathway activity scores for each donor by brain-region and cell-type to allow for increased sensitivity in detecting even subtle expression changes.
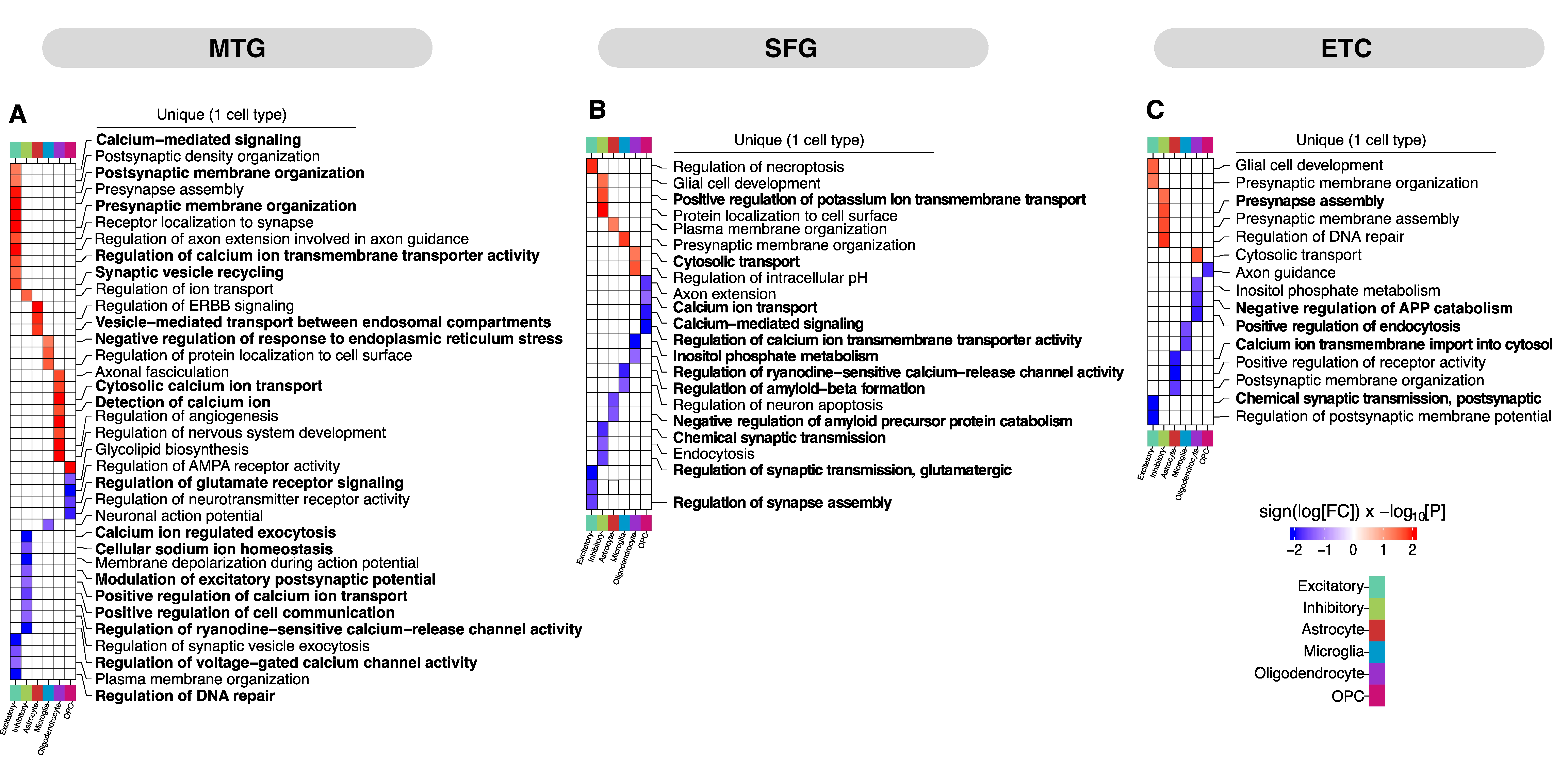
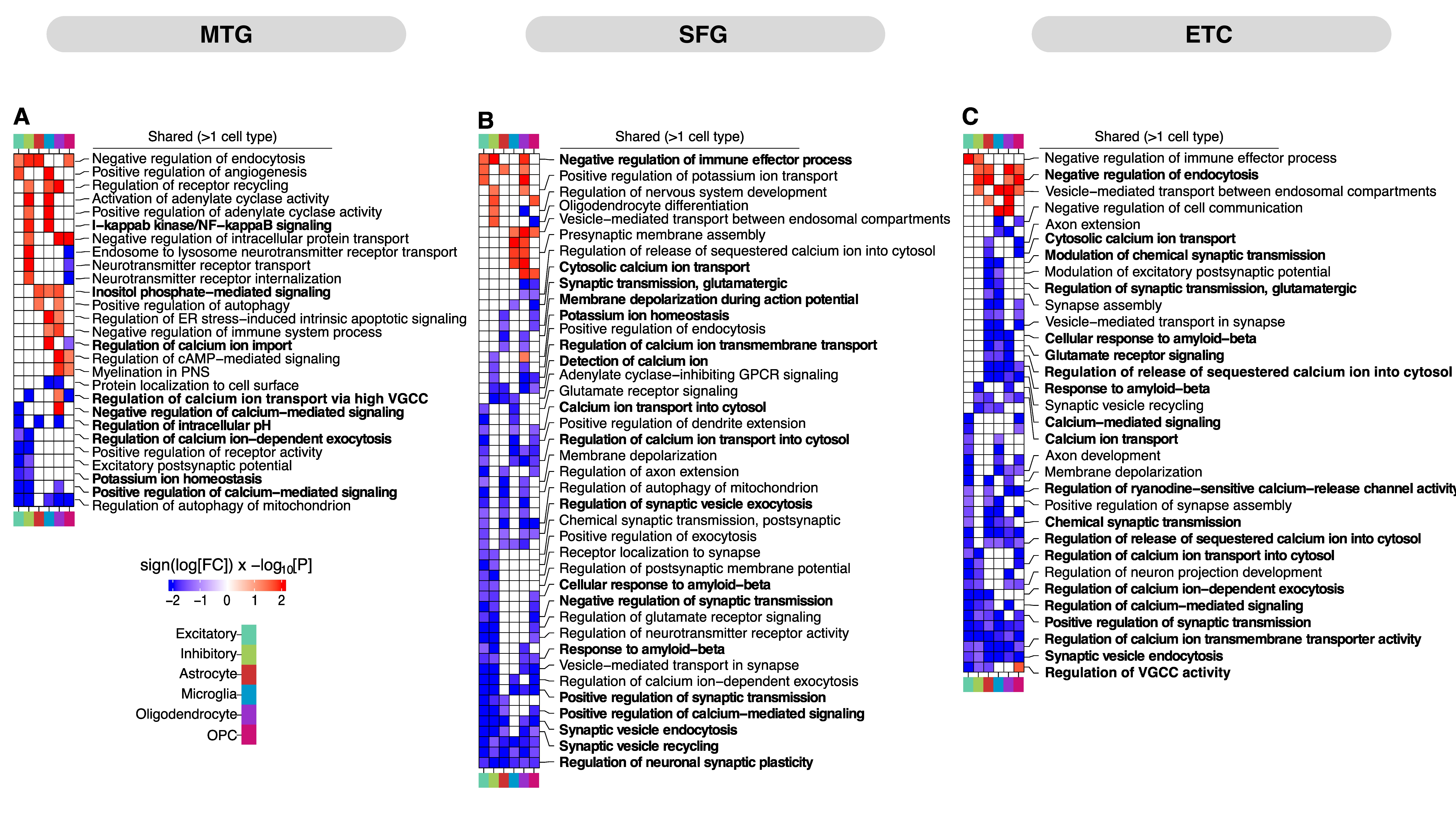
Our analysis revealed that AD inflicts widespread perturbations across molecular processes in the three brain regions, ranging from cell-type-specific alterations to broadly dysregulated pathways affecting multiple cell types. This further highlighted notion of heterogeneity inherent in cellular responses to AD pathogenesis. For instance, in the MTG, neurons showed distinct alterations in synaptic processes like membrane organization, vesicle recycling, and Ca2+ signaling, while glia predominantly exhibited altered vesicle transport, inflammation, and signaling.
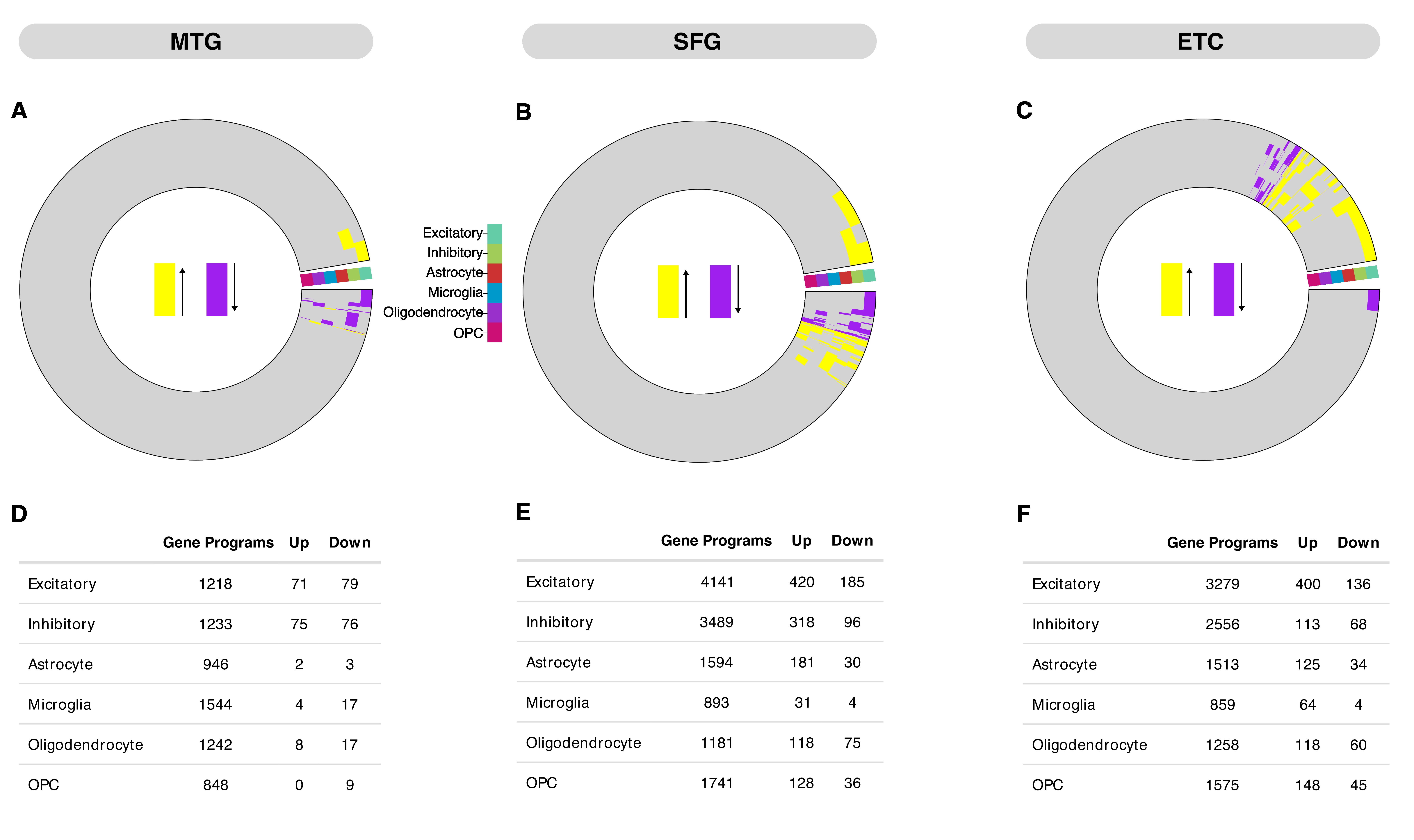
Surprisingly, we found that only a small proportion of differentially expressed genes (DEGs) were associated with the gene programs comprising the previously identified perturbed pathways. This limited, yet dynamic DEG representation indicates that differential gene expression analysis alone does not capture or explain the complexity of AD molecular changes.
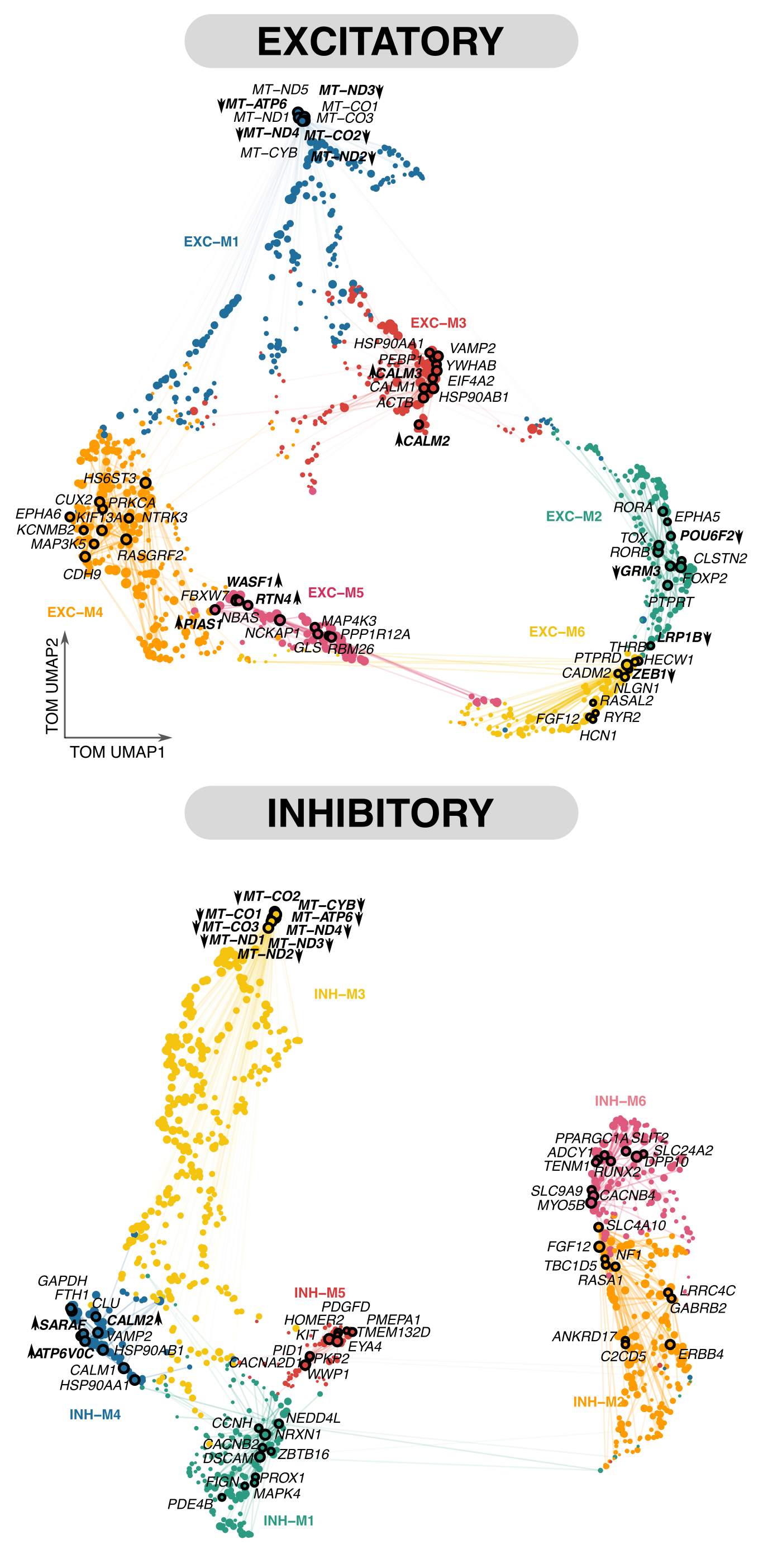

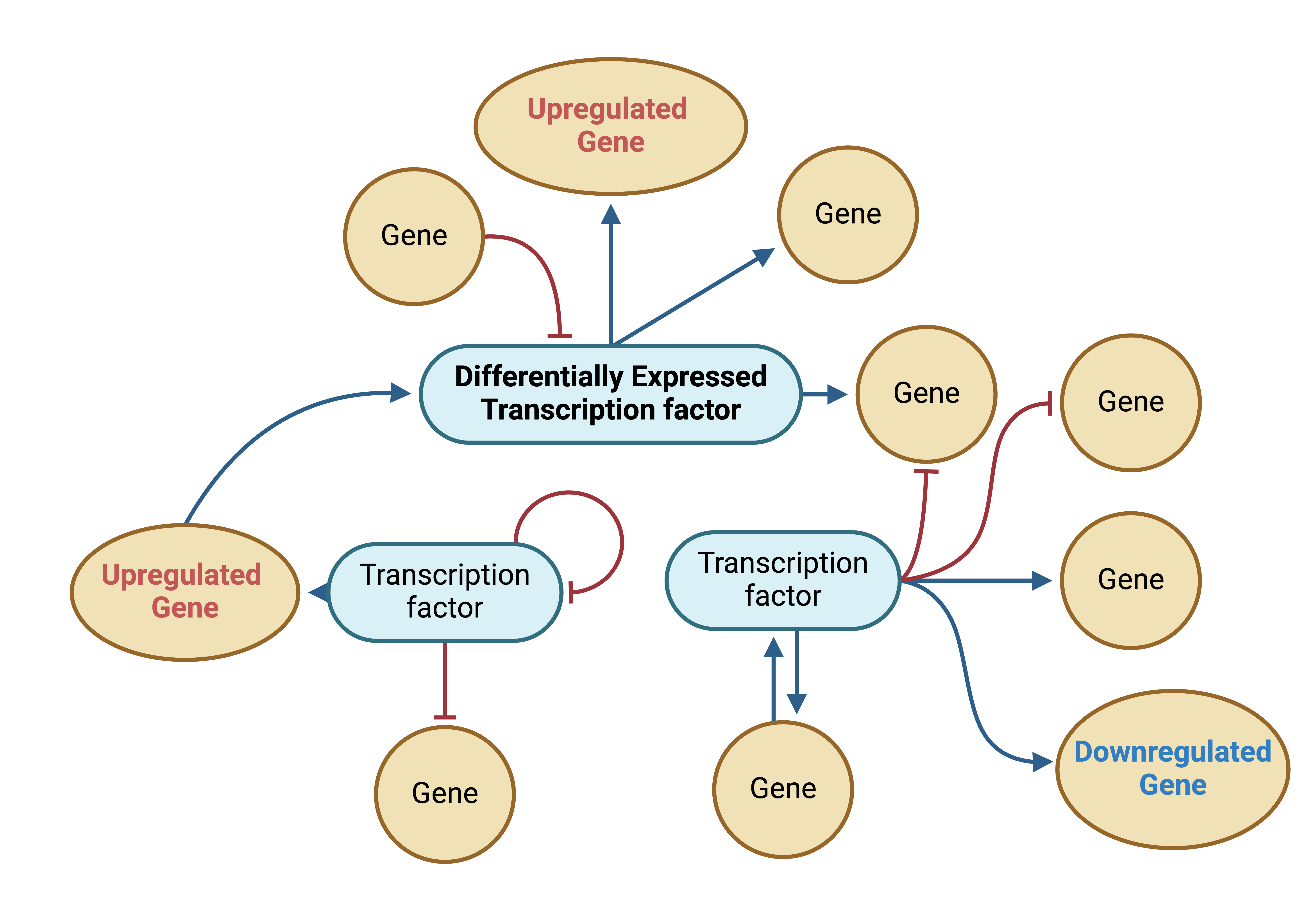
To understand how DEGs influence systems-level dysregulation, we examined gene co-expression networks for each cell type per brain region, focusing specifically on genes within the perturbed pathways. We employed high-dimensional Weighted Gene Co-expression Analysis (hdWGCNA), a framework tailored to handle the sparsity in single-cell transcriptomics data. As a result, we uncovered We identified distinct co-expression modules for each cell-type ithin each brain region, along with hub genes likely playing pivotal regulatory roles due to their extensive module interactions. For instance, in the MTG, we found differentially expressed hub genes (hub-DEGs) like MT-CO1, CALM2,WASF1, concentrated in specific modules, which indicates dysregulation of key pathways like mitochondrial function, Ca2+ signaling, and cytoskeletal dynamics. Surprisngly, only a few glial modules displayed enrichment for hub-DEGs, suggesting a more limited role of DEGs in orchestrating pathway perturbations in glial cells. However, several modules contained AD-related hub genes such as, APOE, CLU, CST3, CD81, and Solute Carrier (SLC) family genes, associated with critical glial functions, such as glutamate regulation, synaptic plasticity, and microglial activation states.
Further analysis of recurrent hub genes across brain regions suggested potential conserved genes driving the observed pathway perturbations in each cell type. For example, excitatory neurons showed substantial overlap in hub genes like ACTB, CALM1/2, GAPDH, HSP90AB1, and UCHL1, participating in critical processes like Ca2+ regulation, autophagy, and synaptic function. Most overlapping hub genes were non-DEGs that co-expressed in similar modules across regions, reinforcing the notion that coordinated network-level dysregulation, rather than changes in individual genes, mediates pathway perturbations.
2. Mapping glia-neuron signaling mechanisms propagating Alzheimer’s disease-associated pathway dysregulations.
Our current work builds on the findings outline above to take a deeper look into the complex interactions between glial cells and neurons in AD. Here, we aim to map the glia-neuron signaling mechanisms that propagate AD-associated pathway dysregulations, with a particular focus on astrocyte-glutamatergic neuron signaling at tripartite synapses.
This work arises due to emerging evidence demonstrating the critical roles of non-neuronal cells, particularly astrocytes and microglia, in the pathological mechanisms underlying nueronal dysfunction in AD progression. Our previous work (outlined above) revealed that in AD, these glial cells exhibit reactive signatures along with significant perturbations of associated regulatory programs. These perturbations have the potential to propagate molecular alterations from glia to neurons, and as a result, contribute to synaptic dysfunction and neuronal loss. However, the precise glia-neuron mechanisms underlying these disrupted signaling processes remain poorly defined, and the extent to which they represent cell-autonomous versus non-cell autonomous effects, remain poorly defined.
To address this gap, we will apply three complementary systems-level strategies which crucially include scRNA-seq and scATAC-seq that cannot be further discussed here to avoid being scooped. Nonetheless, we’re positive our approach will allow us to delineate whether pathway disruptions emerge from cell-autonomous mechanisms (originating within the affected cell type) or non-cell autonomous effects (resulting from dysfunctional signaling between cell types). For instance, we anticipate finding examples of how glial signaling disruptions trigger downstream pathway dysregulation and synaptic dysfunction in neurons during AD progression, representing non-cell autonomous effects. These might include altered glial activities, such as the expression of C1q proteins in disease-associated microglia and disrupted release of thrombospondins from astrocytes, which could initiate a cascade of synaptic pruning and excitotoxic processes through interactions with neuronal receptors.
We will then build on this to focus specifically on disrupted signaling at tripartite synapses between astrocytes and glutamatergic neurons (an area of interest in the lab, for it’s relevance in disorders like seizures and ischemia). Astrocytes play a pivotal role in regulating excitatory glutamate signaling and preventing excitotoxicity, but the precise molecular mechanisms enabling this regulation and how they become dysfunctional in AD are not sufficiently defined. Hence, to address this, we will analyze spatial transcriptomic data from AD and control postmortem tissue across multiple brain regions. This will enable high-resolution spatial mapping of DEGs, ligand-receptor pairs, and dysregulated pathways specifically at tripartite synapse interaction sites. We hypothesize that AD induces pathological changes in critical astrocyte receptors, transporters, and channels that disrupt molecular coordination with neighboring glutamatergic neurons. Ultimately, integrating evidence on localized dysfunctional signaling interactions from spatial omics with results from cell-cell communication from sequencing data, will provide a more robust understanding of the disrupted signaling between astrocytes and neurons that drive excitotoxicity in AD. ***